News and Changelogs
We have increased the default upload cap from 20 to 50 genome sequences. In case you still need larger analyses, please refer to the TYGS FAQ. Please note, the TYGS uses a computing infrastructure to process your request but computing resources are finite and have to be shared among many users across the world. That is, please try to avoid redundant analyses, i.e., please avoid multiple analyses that basically use the same dataset. In case you are unsure on how to use the TYGS, please use the feedback form.
Our new paper describing the relationship between the databases of the Type (Strain) Genome Server (TYGS) and the List of Prokaryotic names with Standing in Nomenclature (LPSN) was recently published in Nucleic Acids Research:
As you might have noticed, the TYGS will remove old job results mainly due to data privacy reasons. The old job result lifetime was 2 weeks and was now extended to 4 weeks. Apart from that, whenever you need your job results a bit longer, just use the contact form to send us a short request and we will extend the lifetime of your job.
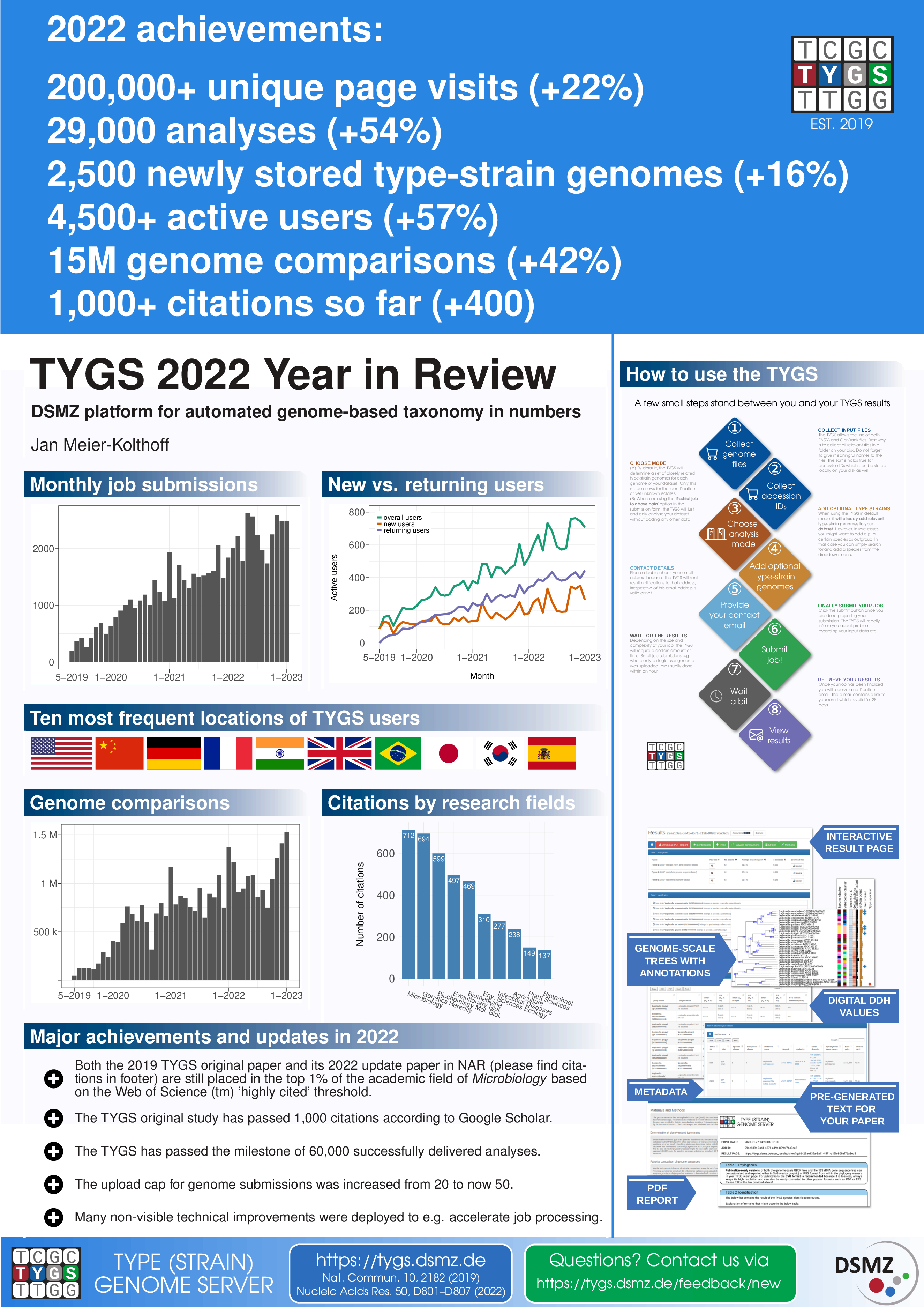
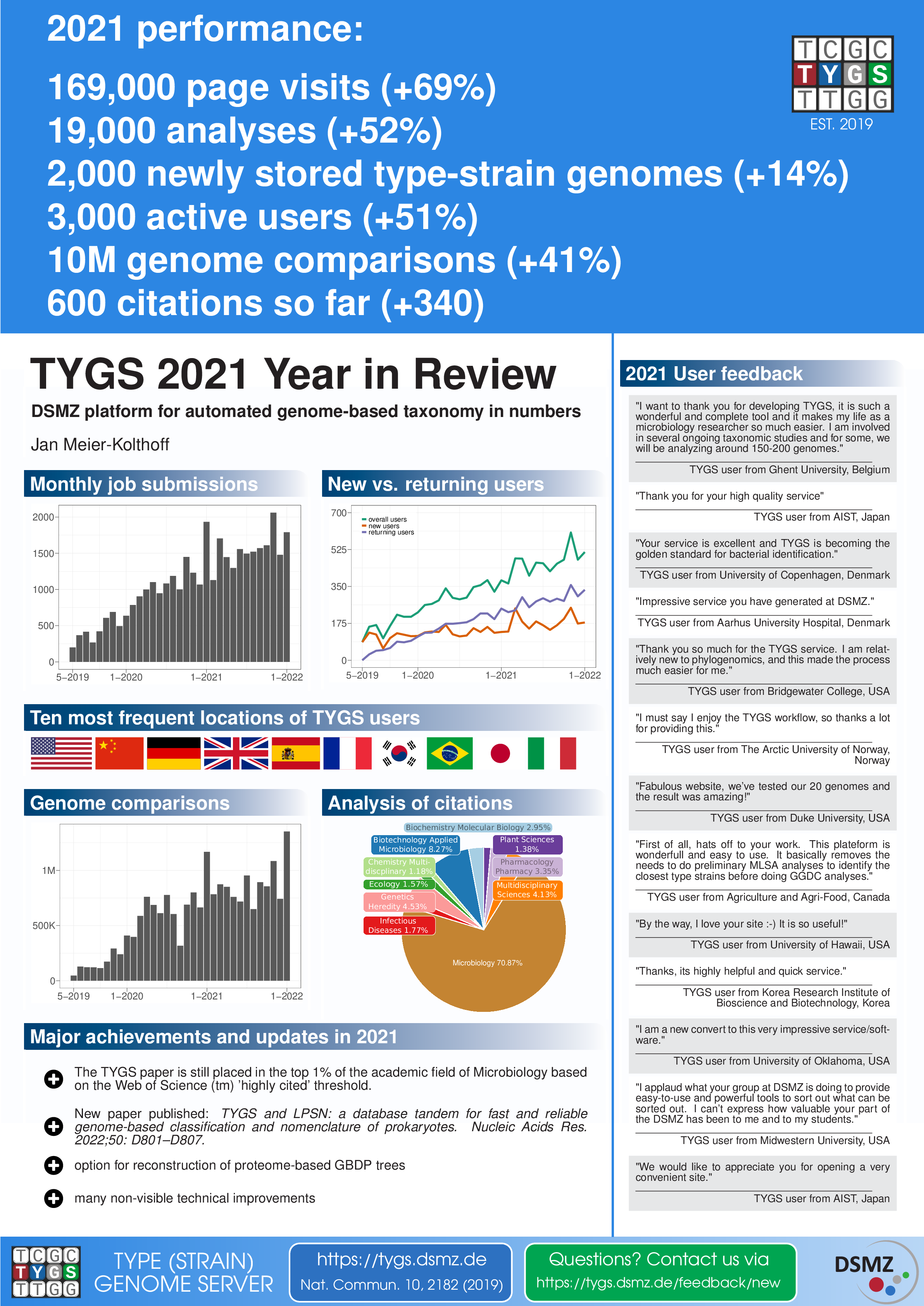
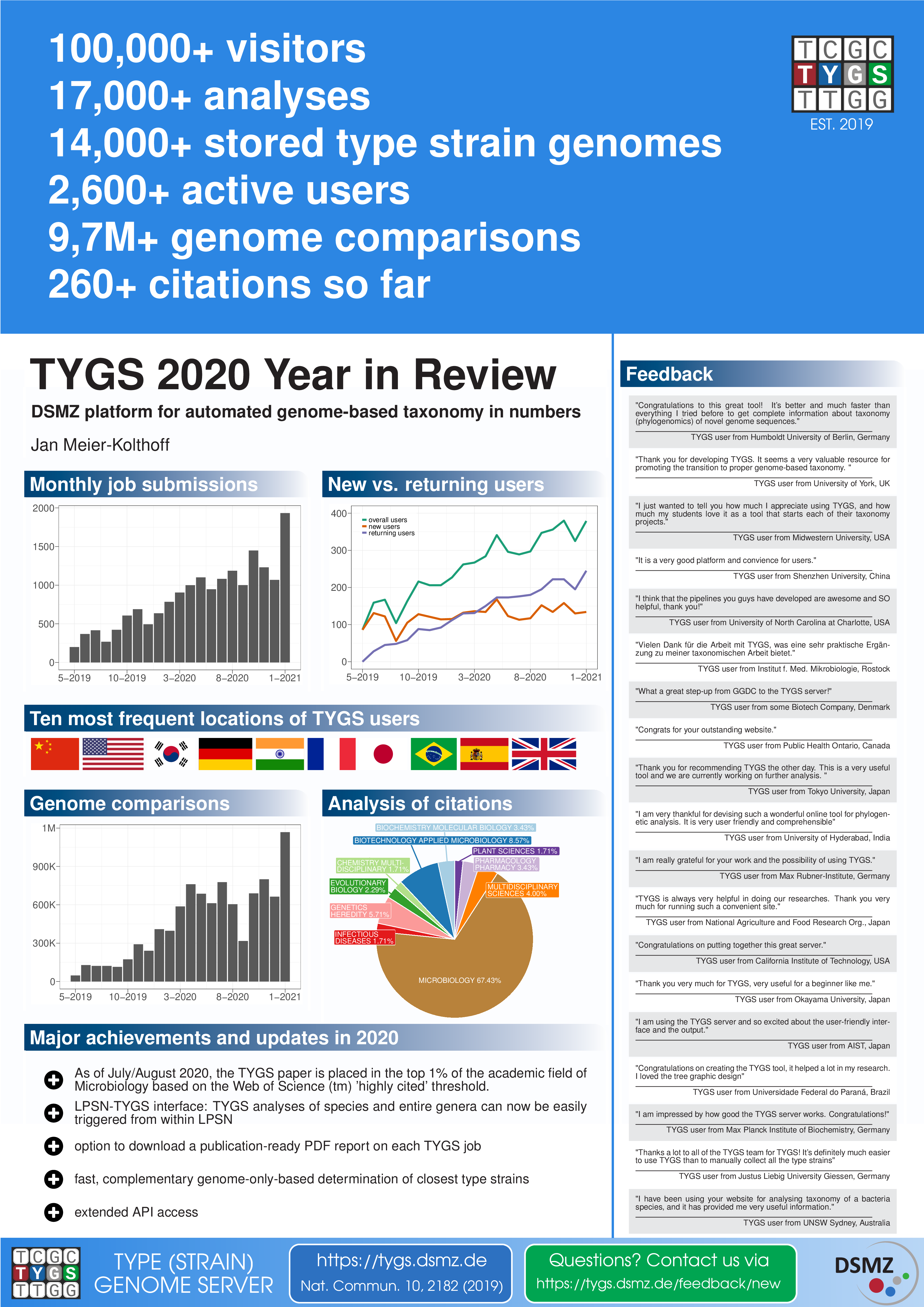
| ~7,000,000 | total number of pairwise genome comparisons |
| ~600,000 | average number of pairwise genome comparisons per month |
| >13,000 | total number of TYGS analyses so far |
| >2,000 | total number of distinct scientists from all around the world so far |
| ~300 | average number of distinct scientists per month |
| ~130 | average number of first-time users per month |